A paper recently caught my eye for two reasons: (1) it implemented an idea that someone smart recently brought up to me and (2) it perfectly illustrates the question of "functional resolution" that comes up in the functional analysis of metagenomes.
The paper is called GraftM: a tool for scalable, phylogenetically informed classification of genes within metagenomes, by Joel A. Boyd, Ben J. Woodcroft and GeneW. Tyson, published in Nucleic Acids Research.
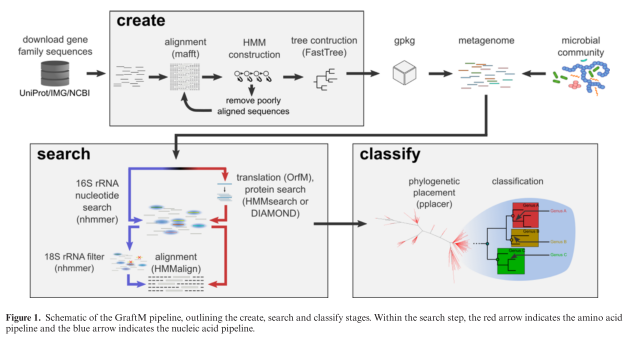
The tool that they present is called GraftM, and it does something very clever (in my opinion). It analyzes DNA sequences from mixtures of microbes (so-called "metagenomes" or microbiome samples) and it identifies the allele of a collection of genes that is present. This is a nice extrapolation of the 16S classification process to include protein-coding sequences, and they even use the software package pplacer, which was developed for this purpose by Erick Matsen (Fred Hutch). Here's their diagram of the process:
They present some nice validation data suggesting that their method works well, which I won't reproduce here except to say that I'm always happy when people present controls.
Now instead of actually diving into the paper, I'm going to ruminate on the concept that this approach raises for the field of functional metagenomics.
Functional Analysis
When we study a collection of microbes, the "functional" approach to analysis considers the actual physical functions that are predicted to be carried out by those microbes. Practically, this means identifying sets of genes that are annotated with one function or another using a database like KEGG or GO. One way to add a level of specificity to functional analysis is to overlay taxonomic information with these measurements. In other words, not only is Function X encoded by this community, we can say that Function X is encoded by Organism Y in this community (while in another community Function X may be encoded by Organism Z). This combination of functional and taxonomic data is output by the popular software package HUMAnN2, for example.
Let's take a step back. I originally told you that protein functions were important (e.g. KEGG or GO labels), and now I'm telling you that protein functions ~ organism taxonomy is important. The difference is one of "functional specificity."
Functional Specificity
At the physical level, microbial communities consist of cells with DNA encoding RNA and producing proteins, with those RNA and protein molecules doing some interesting biology. When we summarize those communities according to the taxa (e.g. species) or functions (e.g. KEGG) that are present, we are taking a bird's eye view of the complexity of the sample. Adding the overlay of taxonomy ~ function (e.g. HUMAnN2) increases the specificity of our analysis considerably.
Now take a look at the figure above. You can see that a single KEGG group actually consists of a whole family of protein sequences which have their own phylogenetic relationships and evolutionary history. This tool (GraftM) presents a means of increasing the level of functional resolution to differentiate between different protein sequences (alleles) that are contained within a single functional grouping (e.g. KEGG label). However, it should be noted that this approach is much more computationally costly than something like HUMAnN2, and is best run on only a handful of protein families at a time.
Summary
Lastly, let me say that there is no reason why a higher or lower level of functional specificity is better or worse. There are advantages at both ends of the spectrum, for computational and analytical reasons that would be tiresome to go into here. Rather, the level of functional specificity should be appropriate to the biological effect that you are studying. Of course, it's impossible to know exactly what will be the best approach a priori, which is why it's great to have a whole collection of tools at your disposal to test your hypothesis in different ways.